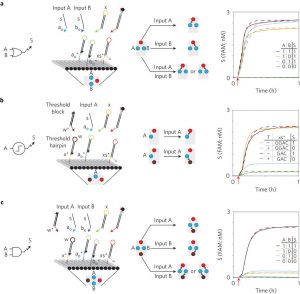
This figure illustrates the elementary logic gates and DNA domino representation. a. Two-input OR. b. Thresholding module. c. Two-input AND. First column: logic gate diagram. Second column: DNA domino representation and top view projection. Credit: Nature Nanotechnology
Spatial organization as a tool for the spread of materials and information is a well-practiced concept. In Mesopotamia, it was used to coax water through a matrix of small channels to water crops. More recently, it has been used to operate semiconductor circuitry. In nature, spatial organization is used by living cells to control the transmission of molecular signals. Researchers at the University of Washington (UW) and scientists at Microsoft Research have used spatial organization to build nanoscale computational circuits made of synthetic DNA.
In a July 24th issue of Nature Nanotechnology, the researchers’ article “A spatially localized architecture for fast and modular DNA computing” discussed this method. Like an AND and OR version of the classic logics game, researchers used “DNA dominoes” as a method of propagating information. In this process, “DNA domino” molecules are positioned at regular intervals on a DNA surface. Information is transmitted when DNA dominoes interact with their immediate neighbors in a cascade.
DNA domino circuits mark an important advance in the field. For decades, researchers have puzzled over the use of DNA molecules for computation. The current components of molecular devices are made from strands of synthetic DNA, where the sequence of the strands determines how they interact. Because billions of DNA molecules rely on a slow process of random diffusion to execute a computation, this method is haphazard in speed and efficiency.
DNA dominoes offer a more robust approach. In this process, DNA dominoes are positioned close to each other on the surface, allowing them to interact quickly with their neighbors. This eliminates the need for random diffusion, leading to an order of magnitude increase in speed. This process also allows for DNA domino recycling. Since their physical location and chemical specificity determines what interactions occur, DNA dominoes can be re-used in multiple locations.
The scaffolding for DNA dominoes was assembled through a process called “DNA origami.” The relatively new process enables unprecedented control over complex molecular structures through the joining of long and short DNA strands. In this process, short strands of DNA (staples) are hinged to a single long strand of DNA (the scaffold).

Georg Seelig
“To build a nanoscale computational circuit, we have to incorporate our individual DNA dominoes into the origami using a special type of extended staple,” PI on the project and UW electrical engineering and computer science and engineering Professor Georg Seelig said. “We do this during the folding process, precisely positioning the DNA molecules on the origami to form a DNA domino.”
The DNA dominoes were assembled into signal transmission lines and elementary Boolean logic gates. By linking the elementary gates together, the researchers were able to create more complex circuits.
This domino approach lays the groundwork for future applications in molecular engineering, offering new opportunities in biosensing, in vitro diagnostics, biomanufacturing and smart therapeutics.
Additional authors on the work include Gourab Chatterjee, a postdoctoral researcher in the UW Department of Bioengineering, Neil Dalchau and Andrew Phillips, researchers at Microsoft Research and Richard Muscat, a former postdoctoral researcher in the UW Department of Electrical Engineering.
—
Information for this article was gathered from a recent Microsoft Research article on the same topic and the paper presented in Nature Nanotechnology.

